DataSpacesAn Extreme Scale Data Management Framework
What is DataSpaces
DataSpaces is a programming system targeted at current large-scale systems and designed to support dynamic interaction and coordination patterns between scientific applications. DataSpaces essentially provides a semantically specialized shared-space abstraction using a set of staging nodes. This abstraction derives from the tuple-space model and can be associatively accessed by the interacting applications of a simulation workflow. DataSpaces also provides services including distributed in-memory associative object store, scalable messaging, as well as runtime mapping and scheduling of online data analysis operations.
DataSpaces is currently being used by production coupled scientific simulation workflow on large-scale supercomputers. For example, as part of the coupled fusion simulation workflow framework, DataSpaces enables memory-to-memory coupling between the gyrokinetic PIC edge simulation code XGC0, and the MHD code M3D-OMP. Similarly, as part of turbulent combustion workflow DataSpaces enables data coupling between the direct numerical simulations (DNS) code S3D and the data analytics pipeline. DataSpaces has been integrated with and deployed as part of the Adaptive IO System (ADIOS2) framework distributed by Oak Ridge National Laboratories. ADIOS2 is an open source I/O middleware package that has been shown to scale to hundreds of thousands of cores and is being used by a very wide range of applications.
Here is a video of DataSpaces demo with ADIOS/Paraview/Pixel3D:
DataSpaces Architecture
DataSpaces has a layered architecture which includes (bottom-up) a communication layer, distributed object store layer, service layer, and programming abstraction layer.
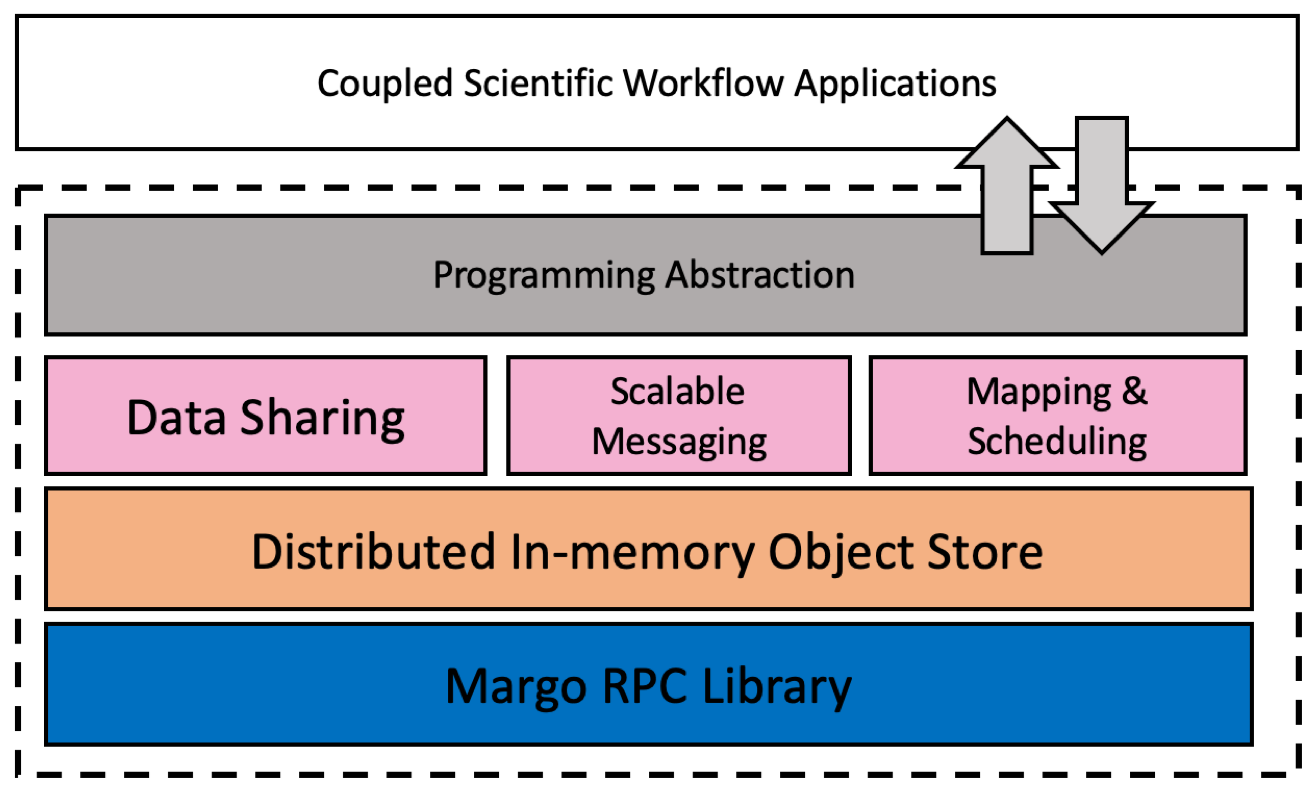 Communication Layer
Communication Layer
DataSpaces uses a high-speed RPC and bulk transfer layer called Margo (mercury + argobots) developed by the Mochi group.
Distributed Object Store Layer
DataSpaces builds an in-memory object storage repository by allocating memory buffers from distributed compute nodes. A distributed hash table (DHT) is constructed to index the data locations and support fast data look-up. Query engine is built on the storage repository and DHT to resolve and service application data queries.
Core Service Layer
On top of the object store and Margo communication layers, DataSpaces implements a number of core services to support the execution and data exchanges in coupled scientific workflows, which are summarized as below.
Coordination and Data Sharing: the service defines and creates an in-memory object storage repository by allocating memory buffers from distributed compute nodes, and manages the memory buffers to create the abstraction of a virtual shared space. The shared-space can be associatively accessed by interacting applications, which would enable asynchronous coordination and memory-to-memory data sharing.
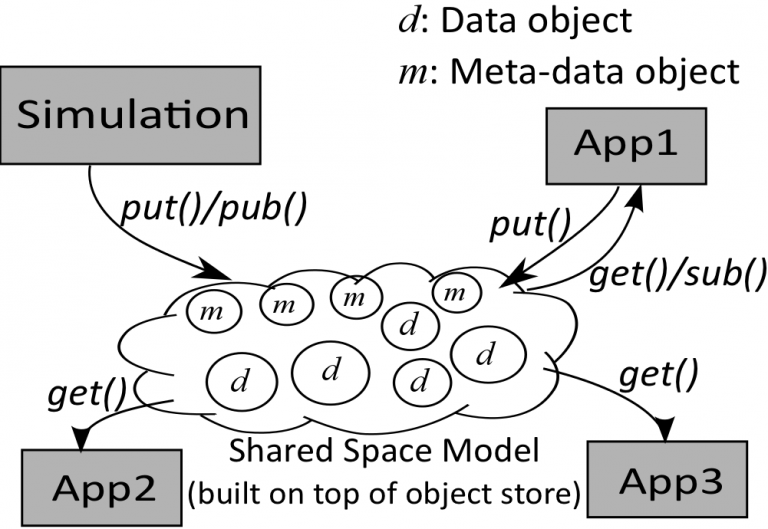
Scalable Messaging: the service enables publish/subscribe/notification type messaging patterns to the scientists. The messaging system allows scientists to (1) dynamically subscribe to data events in regions of interest, (2) define actions that are triggered based on the events, and (3) get notified when these events occur. For example, the registered data event may specify that a function or simple reduction operation of the data values in a certain region of the application domain is greater/less than a threshold value; and the resulting actions include users getting notified and user-defined actions, e.g., visualization or writing the target data to persistent storage, being triggered at the staging nodes.
Mapping and Scheduling: this service manages the in-situ/in-transit placement of online data processing operations as part of the coupled simulation-analytics workflow. In-situ data processing operations execute inline on the same processor cores that run the simulation. In-transit processing executes on the dedicated compute nodes of staging area. The service also supports data-centric mapping and scheduling of the workflow tasks, which aims at increasing the opportunities of intra-node in-memory data sharing and reuse, thus reducing the amount of network data movement.
Programming Abstraction Layer
DataSpaces extends existing parallel programming models, such as MPI and Partitioned Global Address Space (PGAS), with a simple set of APIs and user-defined input files to expose the above mentioned core services, in order to enable the coupling of workflow component applications. (1) DataSpaces provides the put()/get() operators for applications to access the virtual shared-space, thus to enable asynchronous coordination and memory-to-memory data sharing. (2) DataSpaces provides the pub()/sub() operators to enable the publish/subscribe/notification messaging pattern. It allows scientists to dynamically register the data events of interest, define the actions that are triggered based on the events, and get notified when the events occur. (3) Programming data analysis workflow that operate on data being generated by simulation, consists of the following steps: First, define the data dependencies between the analysis operations as a DAG, where each DAG task represents a specific analysis operation of the data analysis workflow; Second, express the data sharing between parent and child tasks of the DAG using shared-space put()/get() operators.